Researchers Produce a Cell Atlas of the Small Intestine
The lining or epithelium of the gut is one of the body’s most diverse and dynamic tissues, an ecosystem of cells that acts as one of the body’s main interfaces with the outside world.
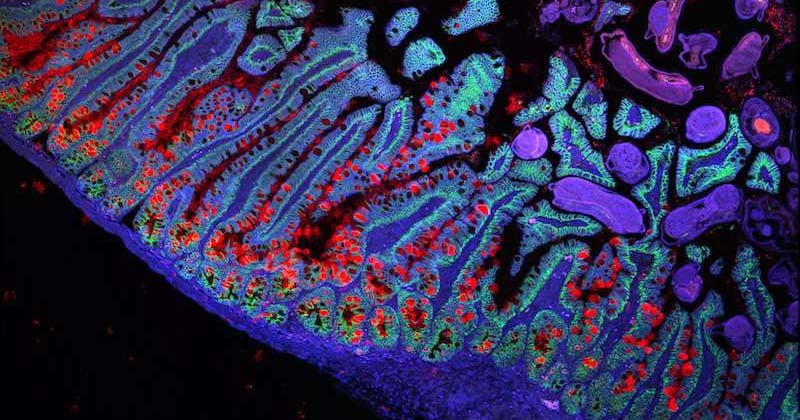
- Adam L. Haber et al. A single-cell survey of the small intestinal epithelium, Nature(2017). DOI: 10.1038/nature24489 Journal reference: Nature*
To better understand this complex tissues and its functions—and the diseases that affect it—a multicenter team led by researchers at the Broad Institute of MIT and Harvard and Massachusetts General Hospital has produced a high-resolution, gene expression-based census of the cells constituting the lining of the small intestine, using more than 53,000 individual cells from the mouse gut or gut organoid models.
This census, published in Nature, comprises a first-draft atlas of the small intestine’s cellular composition, providing a reference for studying the biology of a host of conditions affecting or involving the gut, such as inflammatory bowel disease, cancers of the small intestine, celiac disease, and food allergies. The study also enhances our understanding of the hormones and other signals gut cells produce and sheds new light on how the gut responds to different pathogenic invasions.
The gut has to perform many functions, including absorbing nutrients, generating many of the body’s hormones, and denying entry to noxious substances and pathogens. To do so, it relies on many specialized cells and their specific activities and interactions. Some of these cells are well known, but some have thus far remained unfamiliar.
To carry out their census, the study team relied heavily on single-cell RNA sequencing (scRNA-seq), a suite of genomic techniques capable of identifying specific gene expression profiles within individual cells.
“Every new technology is an opportunity for studying cells and tissues in greater detail,” said Broad core institute member Aviv Regev, a co-corresponding author on the paper, director of the Klarman Cell Observatory (KCO) at the Broad and the institute’s Cell Circuits Program. “They allow us to ask new biological questions or take a fresh look at old ones.
“We wanted to utilize single-cell RNA sequencing to understand what normal intestinal tissue looks like at a deeper level,” she continued. “With that baseline we can start looking at disease.”
“The gut epithelium is in contact with both the immune system and the gut microbiome, and as such the gut is a major hub of cellular connections and therefore it is very important for us to understand gut physiology in health and disease,” said Moshe Biton, a postdoctoral researcher in the KCO and co-first author—with postdoctoral associate Adam Haber and research associate Nogal Rogel—and co-corresponding author on the Nature paper. “It’s also a tissue that we know a great deal about already. So we decided to revisit it and see whether we can find new things using scRNA-seq.”
Leveraging these technologies, the team generated expression profiles for a total of 53,193 small intestinal epithelial cells. Within the data, the team pinpointed expression signatures specific to known cell types (e.g., enterocytes, goblet cells, Paneth cells, tuft cells), specific cell subtypes or populations (e.g., enterocytes at different stages of maturation), and rare cell types (e.g., M cells).
“An exciting outcome of this study was the allocation of known and novel specific sensory molecules associated with each epithelial cell type,” said Rogel. “We hope this will have positive implications in designing drugs targeting metabolic disorders.”
The data also revealed the existence of previously unrecognized cell subtypes and provided support for reclassifying known ones. For instance, the team uncovered a new type of chemical-sensing tuft cell (which helps alert the immune system to infection or other forms of injury) that displayed markers previously thought to be exclusive to immune cells and which may help sound the alarm about allergens and invading parasites.
“We were surprised to see that expression of the gene TSLP—which encodes a cytokine long known to be involved in epithelial-induced inflammation—was exclusive to a particular subset of tuft cells,” Haber noted. “This suggests a significant ‘lookout’ role for these recently characterized cells.”
In addition, the data showed that the gut’s hormone-producing enteroendocrine cells (EECs)—long divided into subsets based on the idea that each only expressed a single hormone—can actually express multiple hormones at once.
The team also mapped their data to different locations along the length of the small intestinal tract. For example, they found that EECs that produce ghrelin (which triggers hunger) tend to group near the beginning of the small intestine (the duodenum, adjacent to the stomach). Those producing peptide YY (which prompts the feeling of being sated), on the other hand, aggregate near the far end (the ileum).
As a proof-of-concept of the atlas’s utility as a reference for disease studies, the researchers also surveyed gut epithelial gene expression in two models of intestinal infection, one of Salmonella bacteria, the other of Heligmosomoides polygyrus, a species of helminth (an intestinal parasite).
Comparisons against the reference census data showed that the intestinal lining dramatically reconfigures itself in response to infection. Certain cell types significantly increased or declined in abundance depending on the kind of infectious insult. Many cells’ expression profiles also changed dramatically, including pathogen-specific changes.
With the reference cell census data in hand, the research team is excited to conduct additional studies, including ones involving models or human patients with gastrointestinal conditions—Crohn’s disease, ulcerative colitis, gastrointestinal cancers, forms of food allergy, to name a few—aimed at identifying changes in gene expression and epithelial structure and function that could reveal new insights and opportunities for therapeutic development.
“We see this atlas providing a foundation for investigating many different questions about the pathology in the gut, effects of drug-induced gut toxicities and for identifying and examining important cells, interactions, and biomarkers,” said Broad institute member Ramnik Xavier, a co-corresponding author on the paper, co-director of the institute’s Infectious Disease and Microbiome Program, and chief of gastroenterology and director of the Center for the Study of Inflammatory Bowel Disease at MGH. “And in creating this resource, we have learned a great deal about how to build an atlas of a complex, dynamic tissue. That kind of knowledge could be important for other efforts focused on other tissues and systems.”













